
|
Running TCW on MacOS |


|
|
|
|||
Problems on Mac
I've run TCW extensively on my Mac, and found a few slight Java problems:- In
runSingleTCW andrunMultiTCW , the interface does not always automatically update until I click in the window. - Pop-up windows can get hidden behind the main window, and it makes TCW appear to be frozen -- move the main TCW window to find the hidden window.
- The buttons are all white.
- The UniProt search links do not work; hence, is not provided on Mac.
Installation on MacOS 10.5.4 (Catalina)
Here are my notes from my installation on my personal Mac v10.15 with Java v14.0.1 (May 2020). It is necessary to use TCW v5.0.3 or higher with MySQL v8.MySQL
- Download from
dev.mysql.com mysql-8.0.17-macos10.14-x86_64.dmg.
Click the '.dmg' file and follow the instructions.
-
There will be a mySQL icon on your System Preferences, where you can start/stop the MySQL server.
- Test: I use tcsh, so I enter the following into the .tcshrc file:
alias mys '/usr/local/mysql/bin/mysql -u root -p[password]'
From the command line, I executesource .tcshrc . If you are using zch or bash, you will need to use their equivalent commands.From the command line, enter 'mys', and then at the prompt:
>create database mytest; >drop database mytest; - In the HOSTS.cfg of the TCW package, set DB_user=[mysql userid], DB_password=[mysql password]
At this point, can build databases and view them.
However, you will not be able to run DE calculations without installing R, or
R and R-Java interface
- Download from cran R the
file R-4.0.0.pkg. Click to install.
- Type 'R' at the terminal prompt, then the following:
>install.packages("rJava") >if (!requireNamespace("biocManager", quietly=TRUE)) >install.packages("BiocManager") >BiocManager::install(version = "3.11") >BiocManager::install("edgeR") >BiocManager::install("goseq") >BiocManager::install("DESeq2") - Set your $R_HOME:
setenv R_HOME /Library/Frameworks/R.framework/Resources # tcsh export R_HOME=/Library/Frameworks/R.framework/Resources # zcsh
- Make sure
runDE has the following:-Djava.library.path=/Library/Frameworks/R.Framework/Resources/library/rJava/jri
Perl-mySQL interface
I could not get this working on Catalina.The following instructions are from an earlier version of this document (for MacOS 10.9.5):
# added 'set $path=($path /usr/local/perl/bin)' to my .tcshrc cpan install DBD::mysql cd .cpan/build/DBD-mysql-4.029-[a unique string will end this directory name] sudo ln -s /usr/local/mysql/lib/*.dylib perl Makefile.PL --testuser=[mysql userid] --testpassword=[mysql password] make make test sudo make install sudo ln -s /usr/local/mysql/lib/libmysqlclient.18.dylib /usr/lib/libmysqlclient.18.dylib
Running /Ext/mac external programs
Since the release of MacOS 10.15, it is impossible to run programs downloaded from the web unless they have been authenticated. Using MacOS Sequoia 15.7.3, I authenticated diamond as follows:- Bring up the
System Setting , selectPrivacy&Security . - Run TCW
Annotate . It will popup a window with 'diamond Not Open'; select Done. - On the
Privacy window, it will now show 'diamond was blocked to protect your Mac.'; select Allow Anyway. - Run TCW
Annotate . It will popup 'Open diamond?'; select Open Anyway. - It will then popup a window asking for your password. Finally, diamond will run.
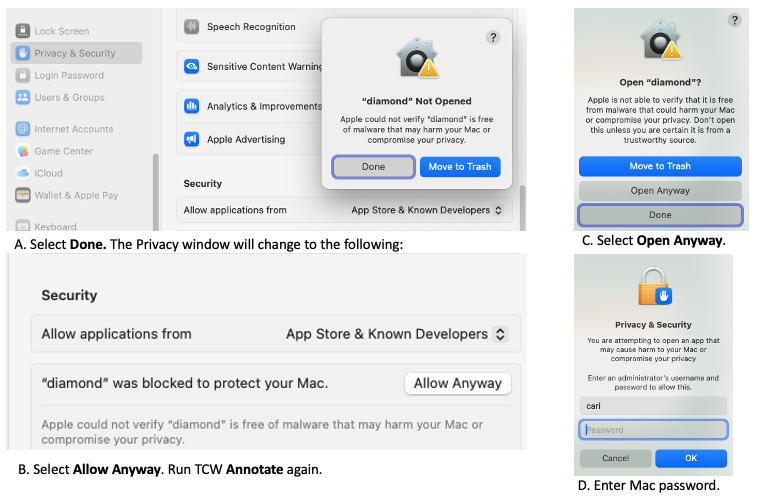
I had to do the same for makeblastdb, blast and tblastx for the
For
bin/
mafft
libexec/
version
countlen
splittbfast
tbfast
disttbfast
dvtditr
f2cl
I could move the /Ext directory around and it would stay authenticated. But when I downloaded a
new TCW_4.tar.gz, I would have to re-authenticate it. Therefore, if you download an updated TCW, use your
previous authenticated /Ext directory.
There may be an easier way to do this, but this works.